Overall objectives
Bioinformatics context: from life data science to functional information about biological systems and unconventional species. Sequence analysis and systems biology both consist in the interpretation of biological information at the molecular level, that concern mainly intra-cellular compounds. Analyzing genome-level information is the main issue of sequence analysis. The ultimate goal here is to build a full catalogue of bio-products together with their functions, and to provide efficient methods to characterize such bio-products in genomic sequences. In regards, contextual physiological information includes all cell events that can be observed when a perturbation is performed over a living system. Analyzing contextual physiological information is the main issue of systems biology.
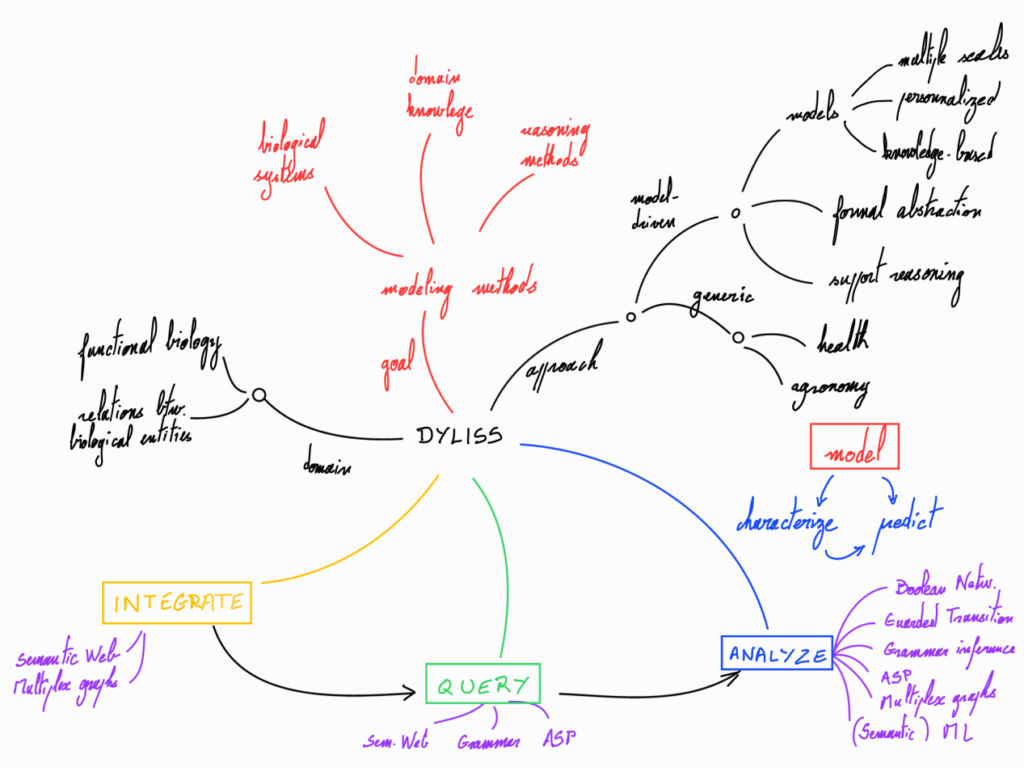
For a long time, computational methods developed within sequence analysis and dynamical modeling had few interplay. However, the emergence and the democratization of new sequencing technologies (NGS, metagenomics) provides information to link systems with genomic sequences. In this research area, the Dyliss team focuses on linking genomic sequence analysis and systems biology. Our main applicative goal in biology is to characterize groups of genetic actors that control the phenotypic response of species when challenged by their environment. Our main computational goals are to develop methods for analyzing the dynamical response of a biological system, modeling and classifying families of gene products with sensitive and expressive languages, and identifying the main actors of a biological system within static interaction maps. We first formalize and integrate in a set of logical or grammatical constraints both generic knowledge information (literature-based regulatory pathways, diversity of molecular functions, DNA patterns associated with molecular mechanisms) and species-specific information (physiological response to perturbations, sequencing…). We then rely on symbolic methods (Semantic Web technologies for data integration, querying as well as for reasoning with bio-ontologies, solving combinatorial optimization problems, formal classification) to compute the main features of the space of admissible models.
Computational challenges. The main challenges we face are data incompleteness and heterogeneity, leading to non-identifiability. Indeed, we have observed that the biological systems that we consider cannot be uniquely identifiable. Indeed, “omics” technologies have allowed the number of measured compounds in a system to increase tremendously. However, it appears that the theoretical number of different experimental measurements required to integrate these compounds in a single discriminative model has increased exponentially with respect to the number of measured compounds. Therefore, according to the current state of knowledge, there is no possibility to explain the data with a single model. Our rationale is that biological systems will still remain non-identifiable for a very long time. In this context, we favor the construction and the study of a space of feasible models or hypotheses, including known constraints and facts on a living system, rather than searching for a single discriminative optimized model. We develop methods allowing a precise and exhaustive investigation of this space of hypotheses. With this strategy, we are in the position of developing experimental strategies to progressively shrink the space of hypotheses and increase the understanding of the system.
Bioinformatics challenges. Our objectives in computer sciences are developed within the team in order to fit with three main bioinformatics challenges (1) data-science and knowledge-science for life sciences (see Section 3.2); (2) understanding metabolism (see Section 3.3); (3) characterizing regulatory and signaling phenotypes (see Section 3.4).
Implementing methods in software and platforms. Seven platforms have been developed in the team during the last five years: Askomics, AuReMe, FinGoc, Caspo, Cadbiom, Logol and Protomata. They aim at guiding the user to progressively reduce the space of models (families of sequences of genes or proteins, families of keys actors involved in a system response or dynamical models) which are compatible with both the knowledge and experimental observations. Most of our platforms are developed with the support of the GenOuest resource and data center hosted in the IRISA laboratory, including their computer facilities [More info]


